
Artificial intelligence, machine learning and deep learning for eye care specialists
Artificial intelligence (AI) methods have been the subject of intense interest in ophthalmology and eye care (1-5). This interest parallels the increasing impact of AI across many domains, including healthcare (6). The availability of machine-learning (ML) frameworks and increasing compute power promise to empower an ever-broader range of people to become AI practitioners (7-9). At the same time, technology employing AI is being assessed in clinical settings, such as diabetic retinopathy screening (10,11).
However, understanding has not kept pace with technological development, including within the eye care community. Many clinicians and researchers do not fully understand the technology, nor its strengths and limitations (12). Thus there is a growing need to understand AI methods, the related concepts of ML and deep learning (DL), and to be able to use that knowledge to rigorously evaluate (as readers or reviewers) publications describing AI technology in eye care.
This review aims to act as a primer on these technologies, with an eye towards the relevance of AI towards ophthalmology applications. The goals of the review are: to explain the important concepts (e.g., AI, ML, and DL) in non-technical terms; to walk through the typical process by which AI technology is currently developed and readied for deployment; to highlight the current state-of-the-art in the deployment of this technology; and to describe both the great potential and possible pitfalls of this technology.
Overview of AI
AI can be broadly defined as technology that produces intelligent behavior, i.e., decision-making behavior that is comparable to that of humans or other animals. However, AI also encompasses a diverse range of technologies, some of which may not closely match popular conceptions of intelligence. For instance, computer vision is a field of AI that aims to develop algorithms to interpret images. Computer vision includes tasks such as identifying objects in “natural” images (e.g., trees in a landscape or cars on a road). However, this process is so intuitive for humans that it may not traditionally be considered “intelligent” behavior, though vision in humans is a complex process from the neurobiology perspective.
Machine learning (ML) is a subset of AI technologies that focuses on developing systems whose performance improves with experience: that is, the system learns from examples instead of being programmed directly. ML systems can take a range of different inputs, and learn an association between those inputs and a desired set of outputs. In ML applications for medicine, inputs may be images [e.g., color fundus photographs, optical coherence tomography (OCT) scans, visual field maps (13,14)], or other types of data such as text-based radiology reports (15,16). Outputs may also vary from diagnoses to the estimation of refractive error, or the risk of future adverse events such as cardiovascular outcomes or disease progression (17-20).
ML systems may be trained using methods that can be broadly categorized as supervised learning or unsupervised learning. Supervised learning involves providing the system with explicit feedback (“supervision”) as to the correct output for every example. For instance, early in the training process, an image classifier may guess at the correct output (e.g., whether an image contains a cat), after which it gets feedback as to whether this guess was correct. By contrast, unsupervised learning involves learning about the structure of data, without requiring knowledge about the correct outputs (21,22). Without the benefit of labels, unsupervised learning tends to require more data than supervised approaches. To benefit from the best of both approaches, a hybrid of both approaches (semi-supervised learning) exists, but has so far been used less commonly in ophthalmology.
ML models employ a range of algorithms to generate predictions1 and learn about data. Much recent interest has focused on the subset of ML called deep learning (DL), which has both supervised and unsupervised forms, and has emerged over the past decade as a powerful approach. DL is in fact a recent variant of an approach that has been in practice since at least the 1950s, often referred to as connectionism (23). This approach involves the construction of artificial neural networks. These networks are comprised of groups of computing units (called “nodes” or “neurons”) which are interconnected in particular arrangements. Most commonly, neural networks are arranged in a hierarchy of sequential “layers”. Nodes in each layer process input data, perform localized processing independently, and output to the next layer. At the top of the hierarchy are output layers, whose nodes represent the model predictions. DL is so named because it uses “deep” networks with many layers in the hierarchy. Though there are many variants of these methods that leverage more complex connection patterns (24-26), the basic principles of connected compute nodes remain true across DL methods.
Developing a deep learning model
The final developed DL system is also termed a “model”. As an illustrative example, we shall consider an example in which a DL model is trained to detect the severity of age-related macular degeneration (AMD; Figure 1). Note that this is an example of a supervised learning task: The network to be trained takes as input pixels in the image and outputs the model’s predicted likelihood for each AMD severity level (none, early, intermediate, advanced dry, advanced wet; Figure 1A).
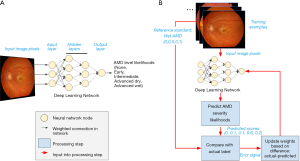
The goals of training the model are: to have these predicted likelihoods reflect the actual likelihoods for each retina image; and to ensure that, once trained, the model will also provide accurate predictions for other images it has not seen before.
A schematic of our example DL model is shown in Figure 1A. The DL model comprises many layers of nodes, starting with an input layer, proceeding through a number of intermediate, hidden layers, and ending with a final output layer which provides the model prediction. In our example, the input nodes take individual pixel values from the input image and perform localized processing, such as comparing the values of nearby pixels. Each node (“neuron”) decides how much to fire (“activate”) by integrating information from all its inputs. The intermediate activations are further processed to nodes in hidden layers, whose activations may reflect the presence of localized image features, such as edges or curvature. As information progresses to higher layers, the visual features which would maximally activate each node become progressively more complex, until they may reflect the presence of pathology relevant to AMD grading. Finally, a classifier node summarizes the evidence for each possible diagnosis to generate the final predicted likelihood of every AMD grade.
So far, we have discussed how neural networks are comprised of connected compute nodes, and how layers of these nodes can process information to describe richer and richer concepts. This arrangement of the nodes—including the number and arrangement of nodes in layers, and the types of functions that perform localized calculations on each feature—is called the neural network architecture. Each calculation may contain parameters; for example, a simple mathematical formula like y=2x+3 has the “architecture” of multiplication and addition, and has two parameters, “2”, and “3”. Generally, a modern neural network can contain millions or billions of these parameters. An important contrasting point between the architecture and the parameters is that while architectures are often pre-specified for a project at the outset, the parameters are learned from the data during the process of training the network.
What actually happens to an artificial neural network as it is trained? In the case of supervised learning, during the training process, the model makes guesses as to an appropriate output and receives feedback based on the correctness of the guess (Figure 1B). The feedback is then used to modify the parameters to reduce the error. By repeating this process thousands to millions of times (spread across many different labeled examples), the network gradually becomes more accurate.
Note that the outputs in this example is a set of predicted likelihoods: Rather than specify a single guess, such as “advanced dry AMD”, the model estimates the likelihood of each category of AMD severity. In the example retina image shown at the top of Figure 1B, the model may provide scores of (0, 0.1, 0.1, 0.6, 0.2), indicating roughly a 10% likelihood of early AMD, 10% intermediate AMD, 60% advanced dry AMD, and 20% advanced wet AMD. Ultimately, those using the model will want to determine how to convert these likelihoods to a single severity level for the given case; however, that process is often a separate step from training, which generally optimizes continuous scores.
Continuing our AMD example, suppose the true grade (or “class” in ML terminology) for the first example is advanced wet AMD. (This might be determined by a panel of retina specialists, or a reading center; see the section titled “What is needed to develop AI technology in eye care?” below.) The training process would represent this true value as a set of values for each severity level: (0, 0, 0, 0, 1) to indicate 100% certainty of being advanced wet AMD and 0% for the other categories.
From this true value and the predicted scores, the model can compute a feedback signal that is then fed back to the AI model, and used to update its parameters. A large feedback signal might indicate that the predictions are way off, and parameters may need to be updated substantially, while a smaller signal might cause finer adjustments to the model. In this example, the model may not have learned enough information to distinguish wet from dry AMD, and the feedback can impact how it makes these classifications for future images. The precise formula for computing the feedback signal varies depending on the model. Though the specifics of the mathematics may differ, similar feedback is used for the predictions of continuous values [e.g., for quantifying volumes of retinal subcompartments from OCT scans; (14)].
The manner by which parameters are adjusted, and in which training proceeds in general, is also determined by hyperparameters: these are choices set by the researcher. By contrast, parameters are learned directly from the data. The neural network architecture is a hyperparameter. Another example of a hyperparameter is the rate at which the model updates its parameters for a given feedback, termed the learning rate. Too fast of a learning rate may cause the network to “overshoot” the optimal parameters, while too slow of a learning rate will take a prohibitively long time to train the network.
The many parameters and hyperparameters of modern neural networks have the immense advantage of enabling AI to recognize highly complex patterns, including visual patterns that are so intuitive to humans. However, this power also increases the potential to fit “too well” to the training dataset and generalize poorly, a phenomenon called overfitting.
To help protect against overfitting, ML practitioners tend to split the dataset used to develop the model into a training set that is used to learn the parameters, and a tuning set2 that is used to select or “tune” hyperparameters. A separate “clinical validation” set is typically used to assess the model’s performance on an independent dataset. Because of the ability of modern neural networks to overfit, neither the training nor the tuning sets should be used to make statements about the model’s performance. More importantly, the final validation dataset should be reserved for the final evaluation of the model, after all decisions affecting the model’s hyperparameters have been made, and the model’s parameters have been finalized.
Evaluating AI model performance
Depending on the nature of predictions an AI model makes, a range of different metrics may be employed for evaluation. In the case of numeric predictions (e.g., age, refractive error), performance may be assessed by the mean error—the average absolute difference between the predicted and actual values in a validation set. Numeric predictions may also be analyzed using the R2, the square of the correlation coefficient between predicted and actual values for each example, which measures the amount of variance in the true values that the predicted values explains [e.g., (17,18)]. Other evaluation methods include the Bland-Altman plot that helps to assess for bias in error at the lower or higher ends of the scale (27).
In the case of AI models that make binary predictions—such as the presence or absence of a specific pathology—performance is often described using sensitivity and specificity (Figure 2A). These metrics report the accuracy of a binary classification with respect to a reference standard, separately for positive cases (sensitivity) and negative cases (specificity). Reporting these metrics separately, rather than overall accuracy, is often preferred, because the patient impact of low sensitivity can be different from that for low specificity; and these tradeoffs depend on the clinical context. For instance, in screening situations, high sensitivity may be preferred, since the cost of missing a candidate may be higher; while when deciding on a surgical intervention, high specificity may be preferred, in order to prevent avoidable surgery (12).
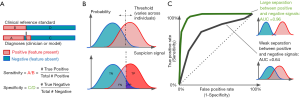
Binary task performance is usually further described using a set of metrics based on the receiver operating characteristic (ROC) curve, which was originally derived from signal detection theory (28,29). This theory aims to quantify how well a signal may be distinguished from non-signal, given that both can be noisy and uncertain.
For instance, consider an ophthalmologist detecting a microaneurysm (MA) on a retina image. These features are visually small, and on some images may look similar to image artifacts; it can be hard to determine with high confidence whether a given feature is an MA. By chance, some features actually caused by artifacts may appear more like an MA; and some actual MAs may appear more like artifacts. How can we quantify how well images from a given camera enable ophthalmologists to detect MAs, given this uncertainty?
Signal detection metrics address this issue by considering how performance varies across a range of possible internal thresholds (Figure 2). Internally, the theory considers an internal response to a given stimulus: This is a continuous value that represents the total evidence for a feature. In our example, an ophthalmologist reviewing a case will have some internal sense of how strongly the image suggests an MA; as the feature looks more like one, this signal goes up. The signal may be higher or lower across different images, but will be higher on average when there actually is an MA, and lower when there isn’t (Figure 2B).
The amount of information available will determine how well-separated the internal responses to positive and negative cases are. A better camera, for instance, may make the difference between MAs and artifacts more distinct. But for a given amount of information, different observers (like doctors) may use different internal thresholds. (Thresholds are also termed “cutoffs”, and “cut-points” in the literature.) For very low thresholds, all examples will be called positive: This implies catching all true cases (100% sensitivity), but overcalling all false cases (0% specificity, or alternately, a 100% false positive rate). As the threshold increases, there will be fewer false positives (increased specificity), but there may start to be more false negatives (decreased sensitivity). When the threshold is very high, all cases would be marked negative: the system will have 100% specificity but 0% sensitivity.
We can quantify the performance across thresholds using an ROC curve3 (Figure 2C). These curves plot the sensitivity against 1-specificity across all thresholds for a given set of internal responses. We show two curves here; in our example, these could represent two different cameras that enable better (green curve) or poorer (black curve) discrimination between MAs and artifact. Each point on an ROC curve is termed an operating point: This represents one particular threshold on an internal response, and a corresponding tradeoff between sensitivity and specificity. The overall performance of each curve is quantified by the area under the curve (AUC). This area ranges from 0 to 1.0. Values around 0.50 indicate chance performance at a task; systems with these values are no better than flipping a coin. Values lower than 0.50 are systematically incorrect. Although there is no absolute threshold for a “good” AUC (it depends critically on the problem), AUCs above 0.7 are considered “acceptable discrimination” (30).
Why now? A historical overview of AI in eye care
The application of computer image analysis to ophthalmological imagery goes back to at least the 1980s (31,32). However, earlier methods tended to focus on more constrained tasks, and have substantially lower documented performance, than recently-developed methods such as DL (4,5,33). For instance, recent DL-based approaches can diagnose a range of diseases using images on a range of patient populations, using a range of cameras [cf. (5,34)].
Early ML applications relied heavily on “hand-crafted” features, which are explicitly determined by the practitioner before training a model. For instance, Algazi et al. (31) analyzed stereo optic disc images, but depended on careful manual alignment of images, along with high-pass filtering and a carefully-specified calculation of expected optic cup depth as assessed by stereo displacement.
By contrast, DL models use as features the raw pixels in the image, and learn what visual features are most relevant to the task, without requiring manual specification of these features. This flexibility has several consequences. It reduces the required manual labor and domain knowledge to apply ML to a task: Practitioners no longer need to specify in advance what information in an example may be diagnostic for an output; they just need a large number of labeled examples. DL can enable higher performance by allowing the model to learn what the discriminative features are. In doing so, DL allows for the possibility of models learning to perform tasks that humans are unable to at present (17,35).
In order to be able to learn on raw features like pixels, DL models typically require abundant compute resources to perform many iterations of learning, as well as a large number of labeled examples to learn from. The former requirement was a more significant constraint prior to the past decade; but access to compute resources has become increasingly available, and this has driven part of the growth in usage of DL. By contrast, access to well-labeled datasets varies across applications. Often, identifying a large and representative set of well-labeled cases for training is the key requirement for developing an ML model for a specific task.
Why ophthalmology is well-positioned for AI technology
Against the backdrop of growing interest in deep learning for medicine, ophthalmology has emerged as a field with a number of important seminal contributions (1-5,7,10,14,17). There are a number of reasons for this. As noted, ML methods and DL in particular, require large datasets with reliable, clinically-relevant labels. This requirement is met in eye care by the growing demand for eye care, particularly for diabetics, and the corresponding rise in “store and forward” teleretinal screening.
Due to the high prevalence of diabetes-related eye conditions such as diabetic retinopathy (DR) and diabetic macular edema (DME) (36,37), large-scale screening programs are being implemented in countries across the globe in order to provide essential eye care for patients at risk of vision loss. This trend is coupled with a lack of access to trained eye care specialists in many of the same regions that have high diabetes prevalence (36,38). As a result of this mismatch between need and availability of care, cloud-based teleophthalmology services have emerged to enable remote grading of images by specialists. This in turn has produced large curated datasets, with labels by experts in the diagnosis of these conditions.
Ultimately, this has enabled ML practitioners to develop high-quality models for detecting DR and DME (4,5,33). Following upon the success of these models, recent efforts have expanded to a range of conditions, including AMD (39-42), glaucoma (5,34,43-45), and retinopathy of prematurity (46-49), among others. At the same time, efforts around AI technologies for DR/DME have begun to progress into real-world settings, with clinical screening programs (11) and a U.S. Food and Drug Administration (FDA) pivotal trial (3).
What is needed to develop AI technology in eye care?
Any AI-powered technology for eye care will need to operate in real-world clinical circumstances. This can place significant constraints to consider when developing these technologies, over and above those driven by technical considerations. We consider both sets of concerns here, with specific attention to cases where technical and real-world considerations interact.
Before implementing an AI model, many inputs are needed. These include, first and foremost, a well-defined clinical problem. What is the existing need the technology will address? It is also useful to separately consider the prediction problem itself—which the AI system will directly provide—and the clinical context in which the prediction is deployed.
For instance, many models have been developed for predicting DR risk (3-5,8). For this same prediction problem, however, there may be distinct clinical problems being solved: The goal may be to increase accuracy of diagnosis in a primary care setting in an automated fashion (3); to assist experts with diagnosis (50); to scale expertise to places with low eye care access, such as remote screening programs (11); or triaging cases which require urgent attention, among others. Understanding up front which task the technology aims to address may affect subsequent decisions around framing of the prediction task, what data to use, and validation approaches.
After determining the clinical problem that the AI model is intended to solve, the steps needed to prepare input data for an AI model are:
- Obtaining a labeled set of examples. These examples should be labeled according to a consistent set of guidelines related to the clinical task; e.g., the International Clinical Diabetic Retinopathy (ICDR) scale for DR (51). It is also important to ensure that the patient population is representative of the actual clinical population, such as based on demographics or disease subtype. Statistical techniques [e.g., (52)] may sometimes be used to improve the precision of estimates in rarer subgroups.
- Partitioning the labeled set into development and validation sets. The development set is often subdivided into training and tune sets, as described in the “Developing a deep learning model” section above. These subsets should be independent at the patient level: no patients used for training or tuning should be used for the final evaluation of the model, or else the model performance may be exaggerated due to memorizing details about particular patients (53).
- Optional, but increasingly common: Obtaining a high-quality reference standard for the validation sets. These reference standard labels are used to provide a more clinically-relevant and rigorous assessment of the model’s performance. Reference standards are specific to each clinical problem. Reference standards may be derived from: standardized reading at established reading centers (3); information from other modalities [e.g., OCT or visual field mapping for models trained on fundus images (34,54)]; or expert adjudication (3,33).
The process by which labeled datasets are obtained can vary dramatically across efforts, and may potentially represent the most significant factor in developing a successful model. For some efforts, public datasets are available for use (55). In other cases, obtaining a labeled dataset requires a large and potentially complex labeling effort. Labeling data can be labor-intensive, both in terms of collecting a large number of labels (4,5), and in some cases collecting a complex set of labels to help assess a complex diagnosis [e.g., glaucoma; (34,45)].
In addition to these inputs, practitioners must make a number of decisions around the development of the ML model:
- An ML implementation framework. For example, many DL models are built using open-source frameworks such as TensorFlow (56), Keras (57), or PyTorch (58);
- A choice of model architecture. In ophthalmology, many successful implementations use standardized, “off-the-shelf” architectures, such as Inception (24,59,60) or ResNet (25). These are architectures that were previously developed and validated on computer vision challenges (61,62);
- Hyperparameters for how the learning process will proceed (see above).
During the implementation, practitioners may check tuning set metrics (such as the AUC; see “Evaluating AI model performance” above) on training and tune sets. At some point, they must determine that a model is “good enough” for final evaluation, reporting, and possible further development. For the iterative training methodology used in most DL approaches, this is typically a point at which the model performance is not increasing substantially for the train or tune sets; and the performance on the train and tune sets is not diverging (which is a sign of overfitting).
Once an AI model has been trained, it should be evaluated on an independent validation set. This typically includes plotting an ROC curve, and reporting the AUC. It may also include identifying an operating point at the ROC curve for which the performance would be useful in a clinical context (12); for instance, if a screening program could accept a 3% false positive rate, the operating point at 97% specificity may be used, and the sensitivity reported (indicating what fraction of positive cases would be successfully detected).
Deploying AI models
Another consideration for a model’s clinical utility is how its performance relates to that of clinicians performing the same task. Papers reporting these models may present the operating points of individual clinicians, or sometimes groups of clinicians, on the same plot as the ROC curve4 (4,5,33,45,64,65). A range of statistical tests, based on contingency tables and/or regression models, have been devised to explicitly test differences in performance between AI models and readers (66-69).
When an AI algorithm has been shown to perform as well or better than clinicians, there is often interest in deploying it into clinical settings. This process is still in its early days: Very few clinical efforts currently use AI methods, with the efforts mostly being used in clinical trials for screening (11,64,70). In the US, there is currently one FDA-approved device for DR screening [IDx-DR; (3)]. The impact of deploying these devices remains to be seen.
Some factors may influence the success of an AI clinical deployment. One factor is the use case: Is the AI intended to create new clinical workflows or improve existing workflows? In some contexts, creating a new automated workflow to screen patients using AI may enable clinicians to direct their attention to the highest risk patients and hence make more efficient use of scarce clinician expertise (6). In other cases, however, it may either be more feasible or impactful, to augment clinicians via AI assistance, while deferring the decision-making to human judgment. Here, the clinical benefit is viewed to be in “up-leveling” performance of clinicians with less training; for instance, AI systems operators at primary care sites (3).
Initial test deployments of AI models have also highlighted some challenges. In a diagnostic study at a primary care practice, Kanagasingam and colleagues (70) found that while an AI-based system correctly identified 2 of 193 patients with referable DR (high sensitivity), it also falsely identified 15 cases of non-referable DR (low specificity), many of which appear to have resulted from low image quality (e.g., dirty lens) and related artifacts. This resulted in a low positive predictive value. It is also possible that selection of a different model operating point may have avoided some of the false referrals. These considerations need to be weighted for future deployments of AI into clinical settings.
What’s next? Future challenges and concerns
While there has been rapid and impressive progress in developing AI systems for eye care, much remains to be done for this technology to fulfill its promise. AI technology has the potential to improve quality of life through more accurate and earlier diagnosis of conditions; through increasing access to eye care; through potentially empowering patients with tools to understand their data; and potentially through other mechanisms not yet elucidated (6). Yet there are many critical steps before this technology is broadly deployed and used in real-world settings.
A central consideration is building trust in AI systems as they are deployed. Without trust in the system, there is a real risk that such systems may be under-utilized, and/or adversely affect diagnostic performance (71). Depending on the method of deployment, many methods may be used to help engender trust. In assisted-read or other human-in-the-loop systems, explanation methods may help clinicians understand and properly utilize predictions from AI systems. These methods provide support for an AI system’s predictions, using different forms of feedback. Many methods focus on attribution of pixels in an image which maximally informed a prediction (72). Other methods, applied in other medical domains, include using structured text data (such as radiology reports) and relating them to images to provide text descriptions supporting a prediction [e.g., describing the likely location and type of a fracture (73)]; or showing images that are determined to be similar to the one a clinician is examining (74). Simpler forms of explanatory support, such as indicating the range of model scores for a multi-class prediction such as DR, may also help clinicians better understand a model’s certainty in a given case; and have been demonstrated to improve performance in simulated settings (50). In addition to explaining a model’s predictions in a specific case, careful training of human operators to use AI tools has been recommended to avoid misapplication of a model’s predictions (75).
As AI systems are deployed, another consideration will be preventing unintended bias in the performance of these systems (76,77). AI systems trained on eye data from one patient population may not necessarily generalize to another population; and systems trained on images taken with one set of equipment may not translate to other equipment. Programs deploying this technology should assess performance across these varying conditions by testing performance on secondary validation sets (datasets that are drawn from a separate population, and/or using different equipment, from the ones the model is trained on). Several recent studies have assessed generalization across patient population and camera type, showing robust generalization (5,64).
Further assessment of bias may be done using newly-developed explanation methods, which can assess how a trained AI system’s predictions may relate to pre-defined concepts (78). This approach, for instance, can determine the extent to which an AI model trained to determine if an image of a person is a doctor is influenced by the gender of the person. Within the ophthalmology domain, this approach has been applied to DR severity diagnoses, confirming that a trained AI model is affected by many of the same concepts that doctors are trained in, such as the presence of MAs, hemorrhages, and neovascularization. Future applications of these methods may help ensure deployed AI systems provide a consistently high standard of care for all patients.
While substantial future efforts will revolve around developing AI for existing indications, other future work may focus on developing new capabilities. In addition to existing work demonstrating the use of AI systems to predict factors like cardiovascular risk (17), refractive error (18), or anemia (35), ophthalmological imagery may predict other signals relevant to a patients’ treatment. Initial work on identifying risk factors for progression of AMD from dry-type to the more severe wet-type (19,20,79) suggests one possible future direction for work of this nature. These new methods hold the potential to extend the range of clinical practice: more than simply performing diagnostic tasks currently performed by clinicians consistently at scale, these newer methods may enable earlier, preventative care or rapid responses to changes in patient disease trajectory.
In conclusion, AI technology holds an exceptional degree of promise across the practice of health care, including eye care. Yet, these are still very much early days. As this review is written, only the very first prospective validations of AI technology into clinical settings have been completed. Notably, assessment of clinical impact in terms of patient outcomes have not yet been performed. Careful examination of these efforts, coupled with thoughtful development of new technologies, can help ensure that the use of AI in ophthalmology is viewed as a highly positive development for eye care providers and patients alike.
Acknowledgments
The authors are grateful to Cameron Chen for helpful comments on the manuscript.
Funding: None.
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editors (Daniel Ting and Haotian Lin) for the series “The State-of-Art Deep Learning Technology in Ophthalmology” published in Annals of Eye Science. The article has undergone external peer review.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/aes.2020.02.05). The series “The State-of-Art Deep Learning Technology in Ophthalmology” was commissioned by the editorial office without any funding or sponsorship. RS reports other from Google LLC/Alphabet, outside the submitted work; In addition, RS has a patent Patents on machine learning application to medical imaging. NH reports personal fees from Google LLC, outside the submitted work; YL reports other from Google LLC, outside the submitted work; In addition, YL has a patent Patents on machine learning for medical images. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
1Though “predictions” classically imply to make statements about the future, this term is used in ML to indicate the ML algorithm making a prediction about input data for which it does not “know” the correct answer.
2Terminology varies across studies. In ML, the datasets are named training, (ML) validation/tuning, and test/holdout set. In the clinical literature (for example derivation of risk scores), there is no tuning dataset, and the datasets are named development/derivation and (clinical) validation.
3The name “receiver operating characteristic” derives from historical reasons; they were originally derived to quantify information broadcast over noisy radio channels. Also due to historical practice, they generally show increasing false positive rate along the X axis; specificity is 100% minus this, so many ROC curves in clinical journals will mark this axis “1-Specificity”.
4One may ask, why are individual clinicians’ performance summarized as single points rather than ROC curves like the model? This depends on whether clinicians are providing explicit judgments as to whether a condition is present, as opposed to a continuous suspicion score. In some studies in radiology, for instance, clinicians provide a value of a continuous suspicion score, which can be used to evaluate a per-clinician ROC curve [e.g., (63)].
References
- Ting DSW, Pasquale LR, Peng L, et al. Artificial intelligence and deep learning in ophthalmology. Br J Ophthalmol 2019;103:167-75. [Crossref] [PubMed]
- Ahuja AS, Halperin LS. Understanding the advent of artificial intelligence in ophthalmology. J Curr Ophthalmol 2019;31:115-7. [Crossref] [PubMed]
- Abràmoff MD, Lavin PT, Birch M, et al. Pivotal trial of an autonomous AI-based diagnostic system for detection of diabetic retinopathy in primary care offices. NPJ Digit Med 2018;1:39. [Crossref] [PubMed]
- Gulshan V, Peng L, Coram M, et al. Development and Validation of a Deep Learning Algorithm for Detection of Diabetic Retinopathy in Retinal Fundus Photographs. JAMA 2016;316:2402-10. [Crossref] [PubMed]
- Ting DSW, Cheung CYL, Lim G, et al. Development and Validation of a Deep Learning System for Diabetic Retinopathy and Related Eye Diseases Using Retinal Images From Multiethnic Populations With Diabetes. JAMA 2017;318:2211-23. [Crossref] [PubMed]
- Topol EJ. High-performance medicine: the convergence of human and artificial intelligence. Nat Med 2019;25:44-56. [Crossref] [PubMed]
- Kern C, Fu DJ, Kortuem K, et al. Implementation of a cloud-based referral platform in ophthalmology: making telemedicine services a reality in eye care. Br J Ophthalmol 2020;104:312-7. [PubMed]
- Rajalakshmi R, Subashini R, Anjana RM, et al. Automated diabetic retinopathy detection in smartphone-based fundus photography using artificial intelligence. Eye 2018;32:1138-44. [Crossref] [PubMed]
- Faes L, Wagner SK, Fu DJ, et al. Automated deep learning design for medical image classification by health-care professionals with no coding experience: a feasibility study. Lancet Digital Health 2019;1:e232-42. [Crossref]
- Ting DSW, Carin L, Abramoff MD. Observations and Lessons Learned From the Artificial Intelligence Studies for Diabetic Retinopathy Screening. JAMA Ophthalmol 2019; [Epub ahead of print]. [Crossref] [PubMed]
- Gulshan V, Rajan RP, Widner K, et al. Performance of a Deep-Learning Algorithm vs Manual Grading for Detecting Diabetic Retinopathy in India. JAMA Ophthalmol 2019; [Epub ahead of print]. [Crossref] [PubMed]
- Shah NH, Milstein A, Bagley PhD SC. Making Machine Learning Models Clinically Useful. JAMA 2019. [Epub ahead of print].
- Elze T, Pasquale LR, Shen LQ, et al. Patterns of functional vision loss in glaucoma determined with archetypal analysis. J R Soc Interface 2015;12:20141118 [Crossref] [PubMed]
- De Fauw J, Ledsam JR, Romera-Paredes B, et al. Clinically applicable deep learning for diagnosis and referral in retinal disease. Nat Med 2018;24:1342-50. [Crossref] [PubMed]
- Lakhani P, Prater AB, Hutson RK, et al. Machine Learning in Radiology: Applications Beyond Image Interpretation. J Am Coll Radiol 2018;15:350-9. [Crossref] [PubMed]
- Zech J, Pain M, Titano J, et al. Natural Language-based Machine Learning Models for the Annotation of Clinical Radiology Reports. Radiology 2018;287:570-80. [Crossref] [PubMed]
- Poplin R, Varadarajan AV, Blumer K, et al. Prediction of cardiovascular risk factors from retinal fundus photographs via deep learning. Nat Biomed Eng 2018;2:158-64. [Crossref] [PubMed]
- Varadarajan AV, Poplin R, Blumer K, et al. Deep Learning for Predicting Refractive Error From Retinal Fundus Images. Invest Ophthalmol Vis Sci 2018;59:2861-8. [Crossref] [PubMed]
- Russakoff DB, Lamin A, Oakley JD, et al. Deep Learning for Prediction of AMD Progression: A Pilot Study. Invest Ophthalmol Vis Sci 2019;60:712-22. [Crossref] [PubMed]
- Arcadu F, Benmansour F, Maunz A, et al. Deep learning algorithm predicts diabetic retinopathy progression in individual patients. npj Digit Med 2019;2:92.
- Nordhausen K. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, Second Edition by Hastie T, Tibshirani R, Friedman J. International Statistical Review 2009; 77: 482–482.
- LeCun Y, Bengio Y, Hinton G. Deep learning. Nature 2015;521:436-444. [Crossref] [PubMed]
- Hebb DO. The Organization of Behavior: A Neuropsychological Theory. New York: John Wiley and Sons 2012.
- Szegedy C, Liu W, Jia Y, et al. Going deeper with convolutions. 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, 2015, pp. 1-9.
- He K, Zhang X, Ren S, et al. Deep Residual Learning for Image Recognition. 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, 2016, pp. 770-778.
- Sutskever I, Vinyals O, Le QV. Sequence to Sequence Learning with Neural Networks. In: Advances in neural information processing systems, pp. 3104-12.
- Altman DG, Bland JM. Measurement in Medicine: The Analysis of Method Comparison Studies. Statistician 1983;32:307. [Crossref]
- Heeger DJ, Landy MS. Signal Detection Theory and Procedures. In: Bruce Goldstein E, (ed). Encyclopedia of Perception. 2010. DOI:
10.4135/9781412972000.n287 . - Hanley JA, McNeil BJ. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 1982;143:29-36. [Crossref] [PubMed]
- Hosmer DW Jr, Lemeshow S, Sturdivant RX. Applied Logistic Regression. Wiley Series in Probability and Statistics 2013. [Epub ahead of print]. DOI:
10.1002/9781118548387 . - Algazi VR, Keltner JL, Johnson CA. Computer analysis of the optic cup in glaucoma. Invest Ophthalmol Vis Sci 1985;26:1759-70. [PubMed]
- Nagin P, Schwartz B, Nanba K. The reproducibility of computerized boundary analysis for measuring optic disc pallor in the normal optic disc. Ophthalmology 1985;92:243-51. [Crossref] [PubMed]
- Krause J, Gulshan V, Rahimy E, et al. Grader Variability and the Importance of Reference Standards for Evaluating Machine Learning Models for Diabetic Retinopathy. Ophthalmology 2018;125:1264-72. [Crossref] [PubMed]
- Liu H, Li L, Wormstone IM, et al. Development and Validation of a Deep Learning System to Detect Glaucomatous Optic Neuropathy Using Fundus Photographs. JAMA Ophthalmol 2019; [Epub ahead of print]. [Crossref] [PubMed]
- Mitani A, Huang A, Venugopalan S, et al. Detection of anaemia from retinal fundus images via deep learning. Nat Biomed Eng 2020;4:18-27. [Crossref] [PubMed]
- Zheng Y, He M, Congdon N. The worldwide epidemic of diabetic retinopathy. Indian J Ophthalmol 2012;60:428-31. [Crossref] [PubMed]
- Yau JWY, Rogers SL, Kawasaki R, et al. Global prevalence and major risk factors of diabetic retinopathy. Diabetes Care 2012;35:556-64. [Crossref] [PubMed]
- Bastawrous A, Hennig BD. The global inverse care law: a distorted map of blindness. Br J Ophthalmol 2012;96:1357-8. [Crossref] [PubMed]
- Grassmann F, Mengelkamp J, Brandl C, et al. A Deep Learning Algorithm for Prediction of Age-Related Eye Disease Study Severity Scale for Age-Related Macular Degeneration from Color Fundus Photography. Ophthalmology 2018;125:1410-20. [Crossref] [PubMed]
- Lee CS, Baughman DM, Lee AY. Deep Learning Is Effective for Classifying Normal versus Age-Related Macular Degeneration OCT Images. Ophthalmology Retina 2017;1:322-7. [Crossref] [PubMed]
- Peng Y, Dharssi S, Chen Q, et al. DeepSeeNet: A Deep Learning Model for Automated Classification of Patient-based Age-related Macular Degeneration Severity from Color Fundus Photographs. Ophthalmology 2019;126:565-75. [Crossref] [PubMed]
- Burlina PM, Joshi N, Pacheco KD, et al. Use of Deep Learning for Detailed Severity Characterization and Estimation of 5-Year Risk Among Patients With Age-Related Macular Degeneration. JAMA Ophthalmol 2018;136:1359-66. [Crossref] [PubMed]
- Li Z, He Y, Keel S, et al. Efficacy of a Deep Learning System for Detecting Glaucomatous Optic Neuropathy Based on Color Fundus Photographs. Ophthalmology 2018;125:1199-206. [Crossref] [PubMed]
- Christopher M, Belghith A, Bowd C, et al. Performance of Deep Learning Architectures and Transfer Learning for Detecting Glaucomatous Optic Neuropathy in Fundus Photographs. Sci Rep 2018;8:16685. [Crossref] [PubMed]
- Phene S, Dunn RC, Hammel N, et al. Deep Learning and Glaucoma Specialists: The Relative Importance of Optic Disc Features to Predict Glaucoma Referral in Fundus Photographs. Ophthalmology 2019;126:1627-39. [Crossref] [PubMed]
- Gupta K, Campbell JP, Taylor S, et al. A Quantitative Severity Scale for Retinopathy of Prematurity Using Deep Learning to Monitor Disease Regression After Treatment. JAMA Ophthalmol 2019; [Epub ahead of print]. [Crossref] [PubMed]
- Redd TK, Campbell JP, Brown JM, et al. Evaluation of a deep learning image assessment system for detecting severe retinopathy of prematurity. Br J Ophthalmol 2018; [Epub ahead of print]. [PubMed]
- Ting DSW, Wu WC, Toth C. Deep learning for retinopathy of prematurity screening. Br J Ophthalmol 2018; [Epub ahead of print]. [PubMed]
- Wang J, Ju R, Chen Y, et al. Automated retinopathy of prematurity screening using deep neural networks. EBioMedicine 2018;35:361-8. [Crossref] [PubMed]
- Sayres R, Taly A, Rahimy E, et al. Using a Deep Learning Algorithm and Integrated Gradients Explanation to Assist Grading for Diabetic Retinopathy. Ophthalmology 2019;126:552-64. [Crossref] [PubMed]
- International Council of Ophthalmology. ICO Guidelines for Diabetic Eye Care. Available online: https://www.idf.org/component/attachments/attachments.html?id=407&task=download (2017, accessed 27 September 2019).
- Mansournia MA, Altman DG. Inverse probability weighting. BMJ 2016;352:i189. [Crossref] [PubMed]
- Zhang C, Bengio S, Hardt M, et al. Understanding deep learning requires rethinking generalization. arXiv preprint arXiv:161103530. Available online: https://arxiv.org/pdf/1611.03530.pdf (2016, accessed 25 September 2019).
- Varadarajan AV, Bavishi P, Raumviboonsuk P, et al. Predicting optical coherence tomography-derived diabetic macular edema grades from fundus photographs using deep learning. Nat Commun 2020;11:130. [Crossref] [PubMed]
- Diabetic Retinopathy Detection. Available online: https://kaggle.com/c/diabetic-retinopathy-detection (accessed 25 September 2019).
- Abadi M, Agarwal A, Barham P, et al. Large-Scale Machine Learning on Heterogeneous Distributed Systems. TensorFlow. Available online: https://www.tensorflow.org/ (2015, accessed 25 September 2019).
- c. Keras. Available online: https://keras.io/ (2015, accessed 25 September 2019).
- Automatic differentiation in PyTorch. Available online: https://openreview.net/pdf?id=BJJsrmfCZ (accessed 25 September 2019).
- Szegedy C, Vanhoucke V, Ioffe S, et al. Rethinking the Inception Architecture for Computer Vision. Available online: http://arxiv.org/abs/1512.00567 (2015, accessed 25 September 2019).
- Szegedy C, Ioffe S, Vanhoucke V, et al. Inception-v4, Inception-ResNet and the Impact of Residual Connections on Learning. Available online: http://arxiv.org/abs/1602.07261 (2016, accessed 25 September 2019).
- Lin TY, Maire M, Belongie S, et al. Microsoft COCO: Common Objects in Context. Available online: http://arxiv.org/abs/1405.0312 (2014, accessed 25 September 2019).
- ImageNet Large Scale Visual Recognition Competition (ILSVRC). Available online: http://image-net.org/challenges/LSVRC/ (accessed 25 September 2019).
- Chan HP, Sahiner B, Helvie MA, et al. Improvement of radiologists’ characterization of mammographic masses by using computer-aided diagnosis: an ROC study. Radiology 1999;212:817-27. [Crossref] [PubMed]
- Raumviboonsuk P, Krause J, Chotcomwongse P, et al. Deep learning versus human graders for classifying diabetic retinopathy severity in a nationwide screening program. NPJ Digit Med 2019;2:25. [Crossref] [PubMed]
- Ardila D, Kiraly AP, Bharadwaj S, et al. End-to-end lung cancer screening with three-dimensional deep learning on low-dose chest computed tomography. Nat Med 2019;25:954-61. [Crossref] [PubMed]
- Yang Z, Sun X, Hardin JW. A note on the tests for clustered matched-pair binary data. Biom J 2010;52:638-52. [Crossref] [PubMed]
- Fagerland MW, Lydersen S, Laake P. Recommended tests and confidence intervals for paired binomial proportions. Stat Med 2014;33:2850-75. [Crossref] [PubMed]
- Obuchowski NA. On the comparison of correlated proportions for clustered data. Stat Med 1998;17:1495-507. [Crossref] [PubMed]
- Liu JP, Hsueh HM, Hsieh E, et al. Tests for equivalence or non-inferiority for paired binary data. Stat Med 2002;21:231-45. [Crossref] [PubMed]
- Kanagasingam Y, Xiao D, Vignarajan J, et al. Evaluation of Artificial Intelligence-Based Grading of Diabetic Retinopathy in Primary Care. JAMA Netw Open 2018;1:e182665 [Crossref] [PubMed]
- Kohli A, Jha S. Why CAD Failed in Mammography. J Am Coll Radiol 2018;15:535-7. [Crossref] [PubMed]
- Gilpin LH, Bau D, Yuan BZ, et al. Explaining Explanations: An Overview of Interpretability of Machine Learning. 2018 IEEE 5th International Conference on Data Science and Advanced Analytics (DSAA). Epub ahead of print 2018. DOI:
10.1109/dsaa.2018.00018 . - Gale W, Oakden-Rayner L, Carneiro G, et al. Producing Radiologist-Quality Reports for Interpretable Deep Learning. 2019 IEEE 16th International Symposium on Biomedical Imaging (ISBI 2019). Epub ahead of print 2019. DOI:
10.1109/isbi.2019.8759236 . - Hegde N, Hipp JD, Liu Y, et al. Similar image search for histopathology: SMILY. NPJ Digit Med 2019;2:56. [Crossref] [PubMed]
- Cai CJ, Winter S, Steiner D, et al. “Hello AI”: Uncovering the Onboarding Needs of Medical Practitioners for Human-AI Collaborative Decision-Making. Available online: https://ai.google/research/pubs/pub48431/ (2019, accessed 24 September 2019).
- Challen R, Denny J, Pitt M, et al. Artificial intelligence, bias and clinical safety. BMJ Qual Saf 2019;28:231-7. [Crossref] [PubMed]
- Zou J, Schiebinger L. AI can be sexist and racist - it’s time to make it fair. Nature 2018;559:324-6. [Crossref] [PubMed]
- Kim B, Wattenberg M, Gilmer J, et al. Interpretability Beyond Feature Attribution: Quantitative Testing with Concept Activation Vectors (TCAV). Available online: http://arxiv.org/abs/1711.11279 (2017, accessed 24 September 2019).
- Babenko B, Balasubramanian S, Blumer KE, et al. Predicting Progression of Age-related Macular Degeneration from Fundus Images using Deep Learning. Available online: http://arxiv.org/abs/1904.05478 (2019, accessed 24 September 2019).
Cite this article as: Sayres R, Hammel N, Liu Y. Artificial intelligence, machine learning and deep learning for eye care specialists. Ann Eye Sci 2020;5:18.


