
In vitro models of retinal diseases
Introduction
Accumulating evidence has progressively highlighted the essential functions of glial cells, which can no longer be thought of as the “silent supporters” of neurons. Within the vertebrate retina, which is responsible for visual perception, Müller cells are the principal glial cells. These cells are radially organized across the entire retinal thickness as originally shown by Cajal through Golgi staining in 1892. This localization is designed for them to play major supporting roles: indeed, these glial cells constitute an anatomo-functional connection between retinal neurons and blood vessels, vitreous body, and subretinal space (1). Müller cells are deeply involved in the generation of the electroretinogram, highlighting their significant role in both retinal physiological and pathological responses. In addition, they are the major producer of retinal vascular endothelial growth factor (VEGF), providing a crucial contribution to retinal angiogenesis during physiological development but also exacerbating pathological neovascularization (2).
In vitro model systems including Müller primary cultures as well as immortalized cell lines, have been developed; these systems represent useful tools to investigate the physiological and pathological events that occur within the retina under different stress conditions.
Methods
Moorfields/Institute of Ophtalmology-Müller 1 (MIO-M1)
Limb et al. described for the first time the isolation protocol of Müller cells from human retina, and they called this spontaneously immortalized cell line MIO-M1 (3). Nowadays, MIO-M1 are widely used to investigate the role of Müller cells in retinal diseases. For example, their response to hyperoxic or hypoxic exposure (when they are cultured in high or low oxygen tension, respectively), can give insights into their involvement in retinopathy of prematurity (ROP). Furthermore, it can be intriguing to analyze MIO-M1 response following stress conditions, as the case of oxidative stress (OS) (when they are cultured with H2O2) or pathological neovascularization (when they are cultured with VEGF). Below, we describe how to manage MIO-M1 cell lines and how to realize the previous mentioned experimental conditions.
Recipes
Complete cell culture medium:
- Dulbecco’s modified Eagle’s medium (DMEM) high glucose (4.5 g/L) (Merck, Darmstadt, Germany, catalog number: D5671);
- 1% Glutamine (Merck, catalog number: G7513);
- 10% Foetal bovine serum (FBS) (Merck, catalog number: F4135);
- 1% penicillin-streptomycin (PS) (Merck, catalog number: P4333).
Starvation medium:
- DMEM high glucose (4.5 g/L);
- 1% Glutamine;
- 0.1% FBS;
- 1% PS.
Culture of MIO-M1
MIO-M1 cells grow at 37 ℃, 5% CO2 in humified conditions. The cells must be seeded in flasks T75 (Sartstedt, Nümbrecht, Germany, catalog number: 83.3911.002) or petri dishes (Sartstedt, catalog number: 83.3902) and maintained until they reach a subconfluent density. When cell density is about 75%, MIO M1 can be expanded or frozen, according to canonical cell culture protocols. In Table 1, MIO-M1cell number at confluence are reported.
Table 1
| Flask/dish type | Cell density at confluence |
|---|---|
| T75 flask | 2.7×106 |
| TC 100 Petri dish | 1.5×106 |
2.7×106 MIO-M1 cells reach 100% confluence in a TC 75 flask. 1.5×106 MIO-M1 cells reach 100% confluence in a TC 100 Petri dish. MIO-M1, Moorfields/Institute of Ophtalmology-Müller 1.
Hyperoxia experiments
Hyperoxic condition should be performed at 75% oxygen and 5% CO2. Normoxic condition (atmospheric oxygen tension and 5% CO2) is used as a control. Experiments in hyperoxia should be performed following a time course, e.g., 24/48/72 h. At least three independent experiments for each condition (normoxia and hyperoxia) must be used to perform statistical analysis. The suggested cells seeding density is reported in Table 2.
Table 2
| Dish type | Number of seeding cells for time course experiments | ||
|---|---|---|---|
| 24 h | 48 h | 72 h | |
| TC 100 mm Petri dish | 1.0×106 | 7.25×105 | 5.89×105 |
| TC 60 mm Petri dish | 4.0×105 | 2.77×105 | 2.25×105 |
MIO-M1 cells number we suggest seeding in TC 100 or 60 mm Petri dishes for each point of the time course experiments. MIO-M1, Moorfields/Institute of Ophtalmology-Müller 1.
- On the first day, seed the suggested number of MIO-M1 cells in complete cell culture medium;
- The following day, remove the medium from the dish;
- Add MIO-M1 starvation medium;
- After 24 h, remove the starvation medium;
- Add complete cell culture medium;
- Incubate the cells in hyperoxic condition, leaving the control cells in normoxia.
After 24/48/72 h proceed with subsequent analyses.
Hypoxia experiments
Hypoxic condition should be performed at 1% oxygen and 5% CO2. Normoxic condition is used as a control (see above). Experiments in hypoxia should be performed following a time course, e.g., 24/48/72 h. At least three independent experiments for each condition (normoxia and hypoxia) must be used to perform statistical analysis. The suggested cells seeding density is reported in Table 2.
- On the first day, seed the suggested number of MIO-M1 cells in complete cell culture medium;
- The next day, remove the medium from the dish;
- Add MIO-M1 starvation medium;
- After 24 h, remove the starvation medium;
- Add complete cell culture medium;
- Incubate the cells in hypoxic condition, leaving the control cells in normoxia.
After 24/48/72 h proceed with subsequent analyses.
Induction of OS
H2O2 is commonly used to induce OS on cultured cells. It has been demonstrated that MIO-M1 cells proliferation/viability and morphology remain unaltered at H2O2 concentration of 100, 200 or 400 µM, while higher concentrations result in dramatic loss of cell viability and alter morphology (4).
- On the first day, seed the suggested number of cells in complete cell culture medium;
- The next day, remove the medium from the dish;
- Add the starvation medium;
- After 24 h, remove the starvation medium;
- Add OS medium;
- After 24 h proceed with subsequent analyses.
To maximize H2O2 effect, treat the cells with 5 µM ML385 (Bio-Techne, Minneapolis, MN, USA, catalog number: 6243), an inhibitor of Nrf2.
Culture with VEGF
To mimic the conditions of pathological angiogenesis, treat MIO-M1 cells with recombinant human VEGF (Merck, St. Louis, MO, USA, catalog number: SRP3182) diluted 1–5 ng/mL in complete growth medium (VEGF medium) as shown in (4).
- On the first day, seed the suggested number of cells in complete cell culture medium;
- The following day, remove the medium;
- Add the starvation medium;
- After 24 h, remove the starvation medium;
- Add VEGF medium;
- After 24 h proceed with subsequent analyses.
Molecular analyses
Once mimicked a given pathological condition (e.g., ROP ischemic phase by means of hyperoxia, ROP neovascularization phase by means of hypoxia, etc.), it is interesting to investigate whether and which molecular pathways are altered compared to untreated controls. Here, we briefly report protocols regarding qPCR, Western Blot analysis, viability assay (e.g., Trypan blue assay and MTT assay), cell cycle analysis and Reactive Oxygen Species (ROS) detection.
qRT-PCR
Perform total RNA extraction using RNeasy Micro Kit (Qiagen, Hilden, Germany, catalog number: 74104). Use Nanodrop™ spectrophotometer (NanoDrop products, Wilmington, CA, USA) for RNA quantification, and random hexamers (Qiagen, QuantiTect® Reverse Transcription Kit, catalog number: 205313) for cDNA synthesis. Use SsoAdvanced Univ SYBR Grn Suprmix (Biorad, CA, USA, catalog number: 1725274) for qRT-PCR reactions. Perform data analysis using the comparative 2–ΔΔCt method for relative quantification (5).
Western blot
Use RIPA Lysis Buffer System (Santa Cruz, Dallas, Texas, USA, catalog number: sc-24948) for protein extraction; assess protein concentration through Micro BCA™ Protein Assay Kit (Thermofisher, Massachusetts, USA, catalog number: 23235). Carry out Western Blot analysis on 15 µg protein using 4–20% Mini-PROTEAN® TGX™ Precast Protein Gels, 10-well (Biorad, catalog number: 4561093) and Trans-Blot Turbo Midi NC Transfer Packs (Biorad, catalog number: 1704159). Use Clarity Western ECL Substrate (Biorad, catalog number: 1705061) to visualize the signal that will be analyzed with Image Lab Software (Biorad).
Cell viability assays
(I) Trypan blue exclusion assay
Detach MIO-M1 cells with trypsin-EDTA. Add 50% Trypan blue (3Z,3'Z)-3,3'-[(3,3'-dimethylbiphenyl-4,4'-diyl)di(1Z)hydrazin-2-yl-1-ylidene]bis(5-amino-4-oxo-3,4-dihydronaphthalene-2,7-disulfonic acid) (Sigma-Aldrich, catalog number: T8154). Perform cell count using TC20 Automated Cell Counter (Biorad, catalog number: 1450102) and Cell Counting Slides Dual-Chamber for TC10™/TC20™ Cell Counter (Biorad, catalog number: 1450011). At least three counts for each sample must be performed for statistical analysis.
(II) MTT assay
Seed MIO-M1 cells onto 96-well flat bottom microplates. When cells are 60–70% confluent, evaluate their viability with Cell Proliferation Kit I MTT assay (Roche, Monza, Italy, catalog number: 11465007001), in accordance with the manufacturer’s protocol. Use Ultra Microplate reader (Bio-Tek Instruments Inc., Winooski, VT, USA) to measure absorbance at 570 nm.
Cell cycle analysis
Seed MIO-M1 cells onto 10 mm plates. Detach the cells with Trypsin-EDTA, fix and stain them in propidium iodide solution, according to the protocol described in reference (6).
ROS determination
(I) ROS determination with the Fluorescent Probe 2’,7’-Dichlorofluorescin diacetate (DCF)
Seed MIO-M1 cells in 96 well-black plates with flat bottom (Sigma-Aldrich, catalog number: 35845) and wash them with PBS 1X, incubate with 15 µm DCF (Sigma-Aldrich, catalog number: 35845) and quantify ROS production following the protocol described in (7).
(II) Flow cytometry analysis of MitoSOX signal
Stain MIO-M1 cells with 1 µm MitoSOX Red (Thermo Fisher Scientific, catalog number: M36008) for 40 min at 37 ℃ following manufacturer’s instructions. After staining, cells can be analyzed by flow cytometry as described in (7).
Immunocytochemistry
MIO-M1 cells can be characterized through immunocytochemistry using the Glutamine Synthetase (GS) as a specific marker for Müller cells (8). In addition, a widely used target to examine pathological-like general responses of these cells following stress is the Glial Fibrillary Acidic Protein (GFAP). GFAP is a cytoskeletal protein used as a marker of reactive gliosis, and abundantly expressed in Müller cells under stress conditions (9). Below, we describe the protocol to perform immunocytochemistry analysis of MIO-M1 cells:
- Plate MIO-M1 cells onto 24×24 slides (Bio Optica, Milano, Italy, catalog number: 09-02424) with complete cell culture medium and allow them to attach;
- Remove medium and wash the slides with PBS 1X;
- Fix the cells by incubating the slides for 20 min at −20 ℃ with cold methanol (Carlo Erba, Milan, Italy, catalog number: 414816) plus 5% Acetic acid glacial Baker analyzed™ (Thermofisher, catalog number: 6052);
- Rinse with PBS 1X three times for 5 min;
- Permeabilize and block into a humidified chamber at RT in 0.1% triton with 5% bovine serum albumin (BSA) fraction V (Merck, catalog number: 10735086001) for 1 h and cover the slides with parafilm (Merck, catalog number: P7668);
- Incubate the slides overnight with the primary antibody in a humidified chamber at 4 ℃ and cover them with parafilm;
- Rinse the slides three times for 5 min with PBS 1X;
- Incubate them for 1 h with the properly conjugated secondary antibody into a dark humidified chamber;
- Rise them three times for 10 min with PBS 1X in the dark;
- Coverslip the slides with the Fluoroshield™ with DAPI mounting medium (Merck, catalog number: slcc8448).
Figure 1A shows MIO-M1 cells stained with anti-GS antibody (Abcam, Cambridge, United Kingdom, catalog number: ab228590) diluted 1:1,000. The secondary antibody is a Goat Anti-Rabbit IgG H&L (Alexa Fluor® 488) (Abcam, catalog number: ab150077).
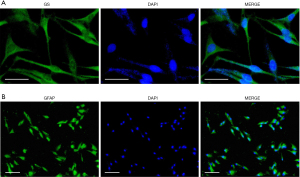
Figure 1B shows MIO-M1 cells stained with monoclonal anti-GFAP antibody (Merck, catalog number: G3893) diluted 1:100, after 24 h hypoxia. The secondary antibody is a Goat Anti-Rabbit IgG H&L (Alexa Fluor® 488) (Abcam, catalog number: ab150077).
Primary cultures of isolated Müller cells
Primary cultures are widely used to investigate the molecular pathways involved in both physiological and pathological conditions, while maintaining the functional properties the cells showed within the organ they come from. Moreover, primary cultures allow us to evaluate the direct effects on cells of either synthetic or natural compounds to be used as drugs in therapy. Particularly, isolation of primary Müller cells offers the invaluable possibility to analyze status and responses of the major retinal glia lineage within diseased retinas. For example, it is possible to isolate Müller cells from diabetic animals, OIR mice, retinal degeneration 10 (RD10) mice, glaucoma rat models and so on. Here, we report the protocol to isolate primary Müller cells from the mouse retina according to (10):
- Dissect mouse retina;
- Incubate the retina in papain and DNase solution (Worthington Papain Kit Worthington Biochemical, catalog number: K003150);
- Cut the retina into small pieces and incubate at 37 ℃ with gentle shaking for 1h until the solution becomes clear;
- Dissociate the mixture with a 5 mL pipette for 2–3 times;
- Centrifuge the cellular suspension at 300 Relative Centrifugal Force (RCF) for five minutes at room temperature;
- Mix the medium (DMEM supplemented with 10% FBS, 1% PS) in a tube with sterile EBSS, reconstituted albumin-ovomucoid inhibitor solution, DNase I solution;
- Remove the supernatant and resuspend the cell pellet in medium mix;
- Add the albumin-inhibitor solution and centrifuge at 70 RCF for 6 minutes at room temperature;
- Remove the supernatant and resuspend the pelleted cells in culture medium;
- Culture the dissociated cells in 0.1% gelatin-coated dish at 5.5% CO2 and 37 ℃;
- Change the medium every 5 days until cells are confluent;
- Pass cells at 1:1 ratio to a fresh dish gelatin-coated dish;
- Adherent Müller glial cells reach more than 95% purity after 2 passages since other retinal neural cells do not survive under such a culture condition.
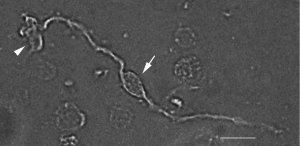
Once isolated, Müller cells can be roughly recognised based on their typical bipolar morphology: they show two main cellular processes, both arising from the central soma (Figure 2). In vivo, their inner process forms an endfoot at the inner limiting membrane which borders the retina to the vitreous humour; instead, their outer process forms close contacts to the photoreceptors (11).
Immunocytochemistry of isolated Müller cells
Once isolated, primary Müller cells can be identified using GS which is expressed throughout their length (8). Below, we report the immunocytochemistry protocol to perform GS staining on isolated Müller cells, according to (12):
- Plate primary Müller cells onto glass slides;
- Remove medium and wash the slides with PBS 1X;
- Fix them for 20 minutes in 4% paraformaldehyde at room temperature (RT) in a humidified chamber;
- Block cells with 3% BSA in PBS 1X for 1 h in a humidified chamber;
- Incubate them with anti-GS antibody overnight at 4 ℃;
- Wash three times with PBS 1X added with 0.1% Tween-20;
- Incubate the slides with the secondary antibody in a dark humidified chamber;
- The nuclei can be stained with 1:1,000 4’,6-diamidino-2-phenylindole (DAPI) for 5 minutes;
- Müller glia can be visualized through a confocal microscope.
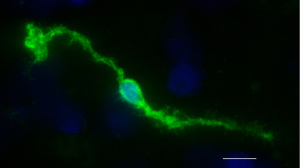
Figure 3 shows immunocytochemistry on primary Müller cells isolated from the mouse retina using anti-GS antibody (Abcam, Cambridge, United Kingdom, catalog number: ab228590) diluted 1:500. The secondary antibody is a Goat Anti-Rabbit IgG H&L (Alexa Fluor® 488) (Abcam, catalog number: ab150077).
Whole-cell patch clamp on isolated Müller cells
Müller cells are deeply involved in controlling osmotic and ionic homeostasis of retinal extracellular environment. To this purpose, they express a variety of ion channels including the glial Inwardly Rectifying K+ (Kir) channels. Kir channels have a high opening likelihood and, thus, a large macroscopic conductance at the resting membrane potential. These channels are open in resting Müller glia and play an important role in the spatial buffering of potassium ions and in retinal osmoregulation (13). Kir 4.1 is the main responsible for the negative resting membrane potential (approximately −80 mV), which is near to the equilibrium potential of K+ ions (14). Several homeostatic functions of Müller cells, as potassium buffering itself and neurotransmitter uptake, depend on this negative membrane potential (1). Müller cells are known to show altered membrane properties in many pathological conditions; whole-cell patch technique is a worthwhile tool to analyze these possible changes in primary Müller cells derived either from diseased animals or treated ones (15). In fact, electrophysiological studies on the retina, particularly on Müller cells, were firstly performed by recording the electroretinogram (ERG) or by using intracellular electrodes or K+-selective electrodes; these methods only allowed to draw indirect conclusion about Müller cells physiology, for example by observing alterations in the ERG following blockage of glial K+ conductance. Instead, whole-cell patch clamp technique enables a more specific investigation of the functionality of channels and receptors on individual cells. This technique was used for investigating Müller cell functions involving electrophysiological properties in different vertebrate species (9). Importantly, whole-cell patch clamp technique can be used to record current signals from different parts of a neuron as well as from a small network of interconnected neurons. Up to date, whole-cell patch clamp recordings have been performed on different neuronal cell types, including retinal ones (16,17).
Below, we report the protocols to perform patch clamp on primary Müller cells isolated from mouse and from rat:
❖ To perform patch clamp recordings from Müller cells isolated from mouse, use borosilicate glass patch pipettes (GB150-8P, Science Products, Hofheim, Germany). Their resistance should be 5–7 MΩ when filled with the intracellular solution reported in (15). To perfuse the recording chamber, use the extracellular solution as described in (15). Record membrane currents of isolated Müller cells in the voltage clamp mode, according to (15);
❖ To perform patch clamp recordings from Müller cells isolated from rats, use a patch clamp amplifier (EPC10; HEKA Electronik, Lambrecht, Germany). The resistance of pipette is 8–10 MΩ when filled with recording solution described in (18). The content of external medium is reported in (18). Kir currents can be evoked through hyperpolarized voltage pulses from a holding potential of −80 mV in increments of 20 mV, according to (18).
Single channel recordings
Single channel recording can be performed on Müller cells in order to specifically examine single-channel properties, as the ones of a Kir4.1 channel, for example. Single-channel patch recordings can be carried out with borosilicate glass pipettes. The content of both the internal solution contained in pipettes, and the bathing solution are reported in (19). Data must be recorded at 5 kHz and low-passed at 1 kHz using an Axon 200B amplifier, according to (19).
Discussions
In summary, both MIO-M1 cells and isolated Müller cells represent valuable in vitro model systems to investigate biological, molecular and electrophysiological properties of a fundamental component of retina. The glial retinal cells, either in healthy or pathological condition. The choice of the model depends on the experimental settings and requirements. Indeed, primary cultures are limited by restricted lifespan since cells, once isolated, undergo senescence and death: no long-lasting studies can be performed on them, differently from continuous MIO-M1 cell line. However, primary Müller cells can be directly isolated from disease animal models, also upon in vivo treatment of the models with drugs to be tested.
Acknowledgments
The authors would like to thank Prof. Dario Rusciano (Sooft Italia, Catania, Italy), Prof. Massimo Dal Monte (University of Pisa, Italy) for introducing us into the interesting topics concerning retinal studies. We are grateful to Dr. Maurizio Cammalleri (University of Pisa, Italy) for his expert assistance in the immunocytochemistry and electrophysiological fields.
Funding: None.
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editor (Dario Rusciano) for the series “Preclinical Models in Ophthalmic Research” published in Annals of Eye Science. The article has undergone external peer review.
Conflicts of Interest: Both authors have completed the ICMJE uniform disclosure form. (available at https://aes.amegroups.com/article/view/10.21037/aes-21-36/coif). The series “Preclinical Models in Ophthalmic Research” was commissioned by the editorial office without any funding or sponsorship. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Reichenbach A, Bringmann A. New functions of Müller cells. Glia 2013;61:651-78. [Crossref] [PubMed]
- Li X, Liu J, Hoh J, et al. Müller cells in pathological retinal angiogenesis. Transl Res 2019;207:96-106. [Crossref] [PubMed]
- Limb GA, Salt TE, Munro PM, et al. In vitro characterization of a spontaneously immortalized human Müller cell line (MIO-M1). Invest Ophthalmol Vis Sci 2002;43:864-9. [PubMed]
- Rossino MG, Lulli M, Amato R, et al. Oxidative Stress Induces a VEGF Autocrine Loop in the Retina: Relevance for Diabetic Retinopathy. Cells 2020;9:1452. [Crossref] [PubMed]
- Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative C(T) method. Nat Protoc 2008;3:1101-8. [Crossref] [PubMed]
- Agnarelli A, Natali M, Garcia-Gil M, et al. Cell-specific pattern of berberine pleiotropic effects on different human cell lines. Sci Rep 2018;8:10599. [Crossref] [PubMed]
- Garcia-Gil M, Turri B, Gabriele M, et al. Protopine/Gemcitabine Combination Induces Cytotoxic or Cytoprotective Effects in Cell Type-Specific and Dose-Dependent Manner on Human Cancer and Normal Cells. Pharmaceuticals (Basel) 2021;14:90. [Crossref] [PubMed]
- Riepe RE, Norenburg MD. Müller cell localisation of glutamine synthetase in rat retina. Nature 1977;268:654-5. [Crossref] [PubMed]
- Grosche J, Härtig W, Reichenbach A. Expression of glial fibrillary acidic protein (GFAP), glutamine synthetase (GS), and Bcl-2 protooncogene protein by Müller (glial) cells in retinal light damage of rats. Neurosci Lett 1995;185:119-22. [Crossref] [PubMed]
- Liu X, Tang L, Liu Y. Mouse Müller Cell Isolation and Culture. Bio Protoc 2017;7:e2429. [Crossref] [PubMed]
- Pannicke T, Ivo Chao T, Reisenhofer M, et al. Comparative electrophysiology of retinal Müller glial cells-A survey on vertebrate species. Glia 2017;65:533-68. [Crossref] [PubMed]
- Wang J, He C, Zhou T, et al. NGF increases VEGF expression and promotes cell proliferation via ERK1/2 and AKT signaling in Müller cells. Mol Vis 2016;22:254-63. [PubMed]
- Newman EA, Frambach DA, Odette LL. Control of extracellular potassium levels by retinal glial cell K+ siphoning. Science 1984;225:1174-5. [Crossref] [PubMed]
- Newman EA. Membrane physiology of retinal glial (Müller) cells. J Neurosci 1985;5:2225-39. [Crossref] [PubMed]
- Reisenhofer M, Pannicke T, Reichenbach A, et al. Characteristics of Müller glial cells in MNU-induced retinal degeneration. Vis Neurosci 2016;33:E013. [Crossref] [PubMed]
- Petrucci C, Resta V, Fieni F, et al. Modulation of potassium current and calcium influx by somatostatin in rod bipolar cells isolated from the rabbit retina via sst2 receptors. Naunyn Schmiedebergs Arch Pharmacol 2001;363:680-94. [Crossref] [PubMed]
- Matsumoto H, Sugio S, Seghers F, et al. Retinal Detachment-Induced Müller Glial Cell Swelling Activates TRPV4 Ion Channels and Triggers Photoreceptor Death at Body Temperature. J Neurosci 2018;38:8745-58. [Crossref] [PubMed]
- Yang Z, Huang P, Liu X, et al. Effect of adenosine and adenosine receptor antagonist on Müller cell potassium channel in Rat chronic ocular hypertension models. Sci Rep 2015;5:11294. [Crossref] [PubMed]
- Ruiz-Ederra J, Zhang H, Verkman AS. Evidence against functional interaction between aquaporin-4 water channels and Kir4.1 potassium channels in retinal Müller cells. J Biol Chem 2007;282:21866-72. [Crossref] [PubMed]
Cite this article as: Lucchesi M, Marracci S. In vitro models of retinal diseases. Ann Eye Sci 2022;7:9.


